A paradigm shift
“From Shape To Substance”
Tumors are now characterized based on their biomarkers signature rather than their tissue of origin or morphology. This leads to a more direct and precise treatment plan.
Discovering unique therapies to treat an
individualˈs cancer, based on the personalized tumor biomarker signature
www.cancer.gov
Comprehensive
Molecular Profiling:
siParadigm has adopted a multi-omics approach to achieve maximum coverage for potentially actionable aberrations that occur at either the genomic (DNA), transcriptomic (RNA), or proteomic cancer pathway levels
We Also Offer Parallel Germline and Somatic Testing With A Team Of In-House Molecular Pathologists, Academic Oncologists and Genetic Counselors Available To Discuss Test Results With You and Your Patient.
Multi-omic Variation
Precision Medicine
Increases Progression-Free Survival
Patients with refractory metastatic cancer who receive genomically guided therapy have improved PFS ratios and longer median PFS compared to patients who do not receive genomically guided therapy.
Percent of Patients (%)
- 100
- 80
- 60
- 40
- 20
- 0
PFS>1.3
Progression-Free Survival (%)
- 100
- 80
- 60
- 40
- 20
- 0
Days from Therapy Initiation
Our testing approach
is clinically geared towards:

Confirming diagnosis:
e.g., androgen receptor and KIT mutation to
confirm the diagnoses of testicular cancer and GIST, respectively.

Disease classification
(e.g., EGFR, ER)

Assessing prognosis
(e.g., BRAF, Ki-67,TP53 and KRAS)

Optimizing treatment selection,
planning, and predicting response to therapy

Identifying
new therapeutic approaches (e.g., PD-L1)

Discovering
clinical trial opportunities (e.g., PDGFR)
| Indication |
Biomarker |
Therapy |
|---|---|---|
|
Non-small cell lung cancer (NSCLC) |
EGFR exon 19 deletions and EGFR exon 21
L858R alterations |
Gilotrit® (afatinib), lressa® (gefitinib), or Tarceva® (erlotinib) |
| EGFR exon 20 T790M alterations |
Tagrisso® (osimertinib) | |
| ALK rearrangements |
Alecensa® (alectinib), XALKori® (crizotinib), or Zykadia® (ceritinib) |
|
| BRAF V600E |
Tafinlar® (dabrafenib) in combination with Mekinist® (trametinib) |
|
| Melanoma |
BRAF V600E | Tafinlar® (dabrafenib) or Zelborat® (vemurafenib) |
| BRAF V600E and V600K |
Mekinist® (trametinib) or
Cotellic® (cobimetinib) in combination with Zelborat® (vemurafenib) |
|
| Breast cancer |
ERBB2 (HER2) amplification |
Herceptin® (trastuzumab),
Kadcyla® (ado-trastuzumab- emtansine), or Perjeta® (pertuzumab) |
| Colorectal cancer |
KRAS wild-type (absence of mutations in codons
12 and 13) |
Erbitux® (cetuximab) |
| KRAS wild-type (absence of mutations in exons
2, 3, and 4) and NRAS wild type (absence of mutations in exons 2, 3, and 4) |
Vectibix® (panitumumab) |
|
| Ovarian cancer |
BRCA 1/2 alterations |
Rubraca® (rucaparib) |
Maximum Precision
with Broad Coverage:
Modern massive parallel sequencing technologies have
given us tremendous capabilities to investigate multiple molecular biomarkers in parallel to reach
results faster on tiny biopsy specimens. Also, multigene panel testing can help identify driver
mutations for which effective drugs may be available. For example, the NCCN makes the following
recommendations for using multi-gene panels in the evaluation of non-small cell lung cancer
(NSCLC): “The NCCN NSCLC Guidelines Panel strongly endorses broader
molecular profiling with the goal of identifying rare driver mutations for which effective drugs
may already be available or to appropriately counsel patients regarding the availability of
clinical trials. Broad molecular profiling is a key component of the improvement of care of
patients with NSCLC.”
Our comprehensive solid tumor panel contains 500+ genes that cover a wide array of alterations, such as:

Single Nucleotide Variants (SNVs), e.g., EGFR T790M.

Insertions & Deletions (InDels), e.g., BRCA1/2 in breast cancer

Copy Number Variants (CNVs), e.g., ERBB2/HER2 amplifications in breast cancer.

Gene Fusions (RNA), e.g., EML4-ALK fusion in NSCLC.

Splice Variants (RNA), e.g., METex14 exon 14 skipping in NSCLC

Microsatellite Instability (MSI), e.g., Lynch syndrome and colon cancer.

Tumor Mutation Burden (TMB), e.g., Mutation per megabyte as a biomarker for immunotherapy.

Somatic and germline Homologous Recombination Deficiency (HRD) and Loss of Heterzygosity (LOH) for assessing eligibility for PARP inhibitors therapy.
HRD
Homologous recombination deficiency
Homologous recombination pathway is a DNA repair pathway that includes BRCA1&2, ATM and several other Loss of Heterozygosity (LOH) genes (including RAD51C, RAD51D, RAD50, BRIP1, BARD1, CHEK2, MRE11A, NBN, PALB2, MLH1, MSH2, MSH6, PMS2, TP53). Mutations of the Homologous Recombination Repair (HRR) gene(s) may result in impaired DNA repair, thereby providing a selective target for cancer cell death (1,2). Poly ADP Ribose Polymerase (PARP) plays an important role in DNA damage repair. Inhibition of the PARP relevant pathway will not result in cell death as the HRR pathway will compensate for the DNA repair process. However, if the HRR pathway is impaired by a mutation in the HRR gene(s), then inhibiting the PARP pathway in the cancer cell will result in cancer cell death(1,2).

FDA Approved PARP inhibitors:
| Indication |
Biomarker |
Therapy |
|---|---|---|
|
Breast cancer |
BRCA1 and BRCA2 genes |
Lynparza® (olaparib)3 |
| Talzenna® (talazoparib)4 |
||
Ovarian Cancer |
BRCA1 and BRCA2
genes |
Lynparza® (olaparib)- treatment/maintenance5 |
| BRAF V600E and V600K |
||
| Pancreatic Cancer |
BRCA1 and BRCA2 genes |
Lynparza® (olaparib)7 |
| Prostate Cancer |
BRCA1&2, ATM, BARD1, BRIP1, CDK12, CHEK1,
CHEK2, FANCL, PALB2, PPP2R2A, RAD51B, RAD51C, RAD51D, RAD54L |
Lynparza® (olaparib)8,9 |
Frequently
Asked Questions:
At siParadigm, we use Next Generation Sequencing (NGS) to test for somatic and germline mutations in the homologous recombination repair (HRR) genes including BRCA1/2, ATM and 42 loss of heterozygosity (LOH) genes. These mutations result in genomic instability in breast, ovarian, pancreatic, or prostate cancers. This helps identify subsets of patients who may be eligible for PARP inhibitor therapy or clinical trials.
A positive somatic or germline result qualifies your patient for PARP inhibitor therapy. There are several drugs that have been FDA-approved based on BRCA1/2 mutations and certain cancers. Many clinical trials are under way to approve these and other drugs for other gene mutations in the HRR pathway as well as other cancers (see the table above).
Studies suggest that PARP inhibitors are indicated in cases of germine and/or somatic mutations in the Homologous Recombination Repair (HRR) genes. That is why testing is recommended for both. Please note that germline testing requires written patient’s consent because of its familial implications.
HRR pathway genes have different predictive values for different cancers. In general, the most predictive are BRCA1&2, and to a lesser extent, ATM. However, other loss of heterozygosity (LOH) genes are also pertinent (including BARD1, BRIP1, CDK12, CHEK1, CHEK2, FANCL, PALB2, PPP2R2A, RAD51B, RAD51C, RAD51D, RAD54L, etc.).
Experience DeepSightTM

Fast TAT/Results

Low QNS (less than 2%)

Flexible Testing Options

Personalized Consultative Comprehensive Reports
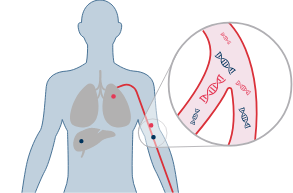
Nucleic Acid from tumors circulates in the blood
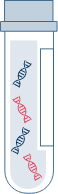
Tumor-associated, cell free nucleic acid is extracted from plasma
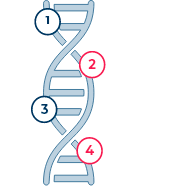
DeepSightTM Liquid Biopsy Assays provide comprehensive coverage of clinically actionable genomic alterations
Liquid Biopsy Assays identify mutational biomarker targets involved in key guidelines and clinical trials for multiple tumor types that are:

Clinically
Actionable

Therapeutically
Predicative

Prognostically
Significant
Beyond the Challenges of Tissue Testing
DeepSightTM Focused and DeepSightTM Comprehensive Guarantee Faster Results and Early Treatment Decisions

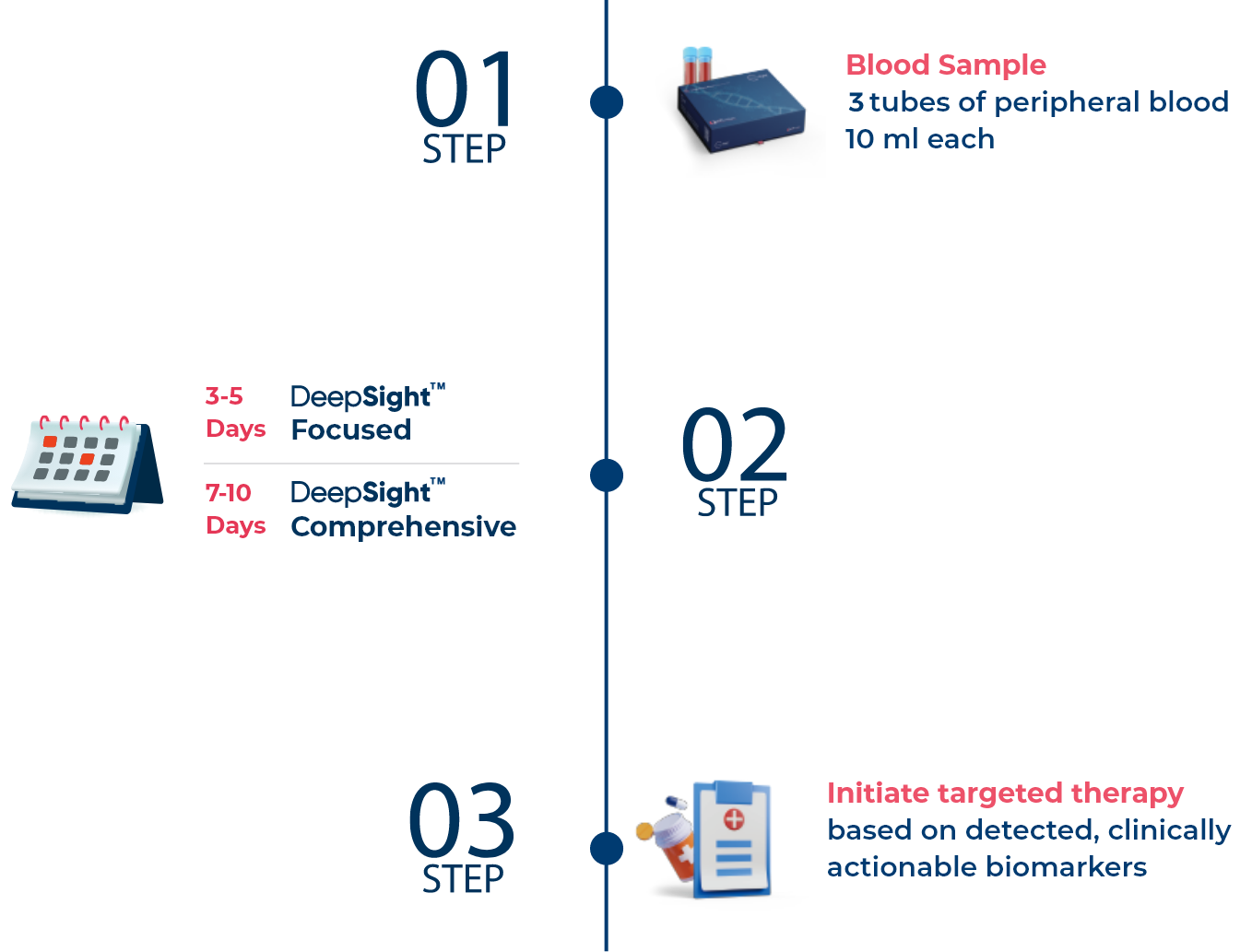

Sample Requirements:
1. Three (3) tubes of peripheral blood (10mL/tube) are collected in PAXgene Blood DNA Tubes. Ship in room temperature (18°C to 25°C) within 24-48 hrs after collection.2.Previously extracted cfDNA is acceptable. A minimum of 50 μL of cfDNA with a concentration of at least 0.60 ng/ μL, corresponding to a total of at least 30ng cfDNA. Optimal cfDNA fragment sizes are between 100-200 bp. Ship in dry ice.

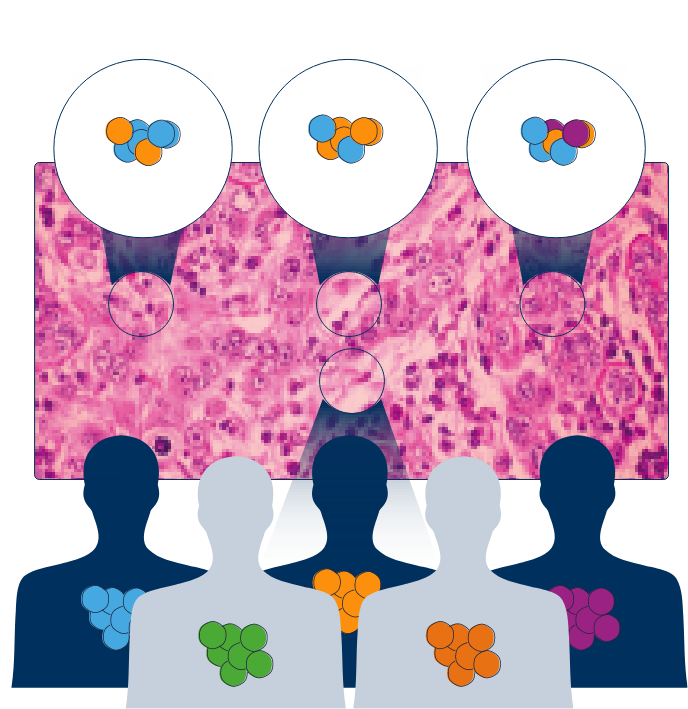
Overcome the dynamic challenges of cancer cells, and capture tumor heterogeneity with our
CONCURRENT TESTING Approach

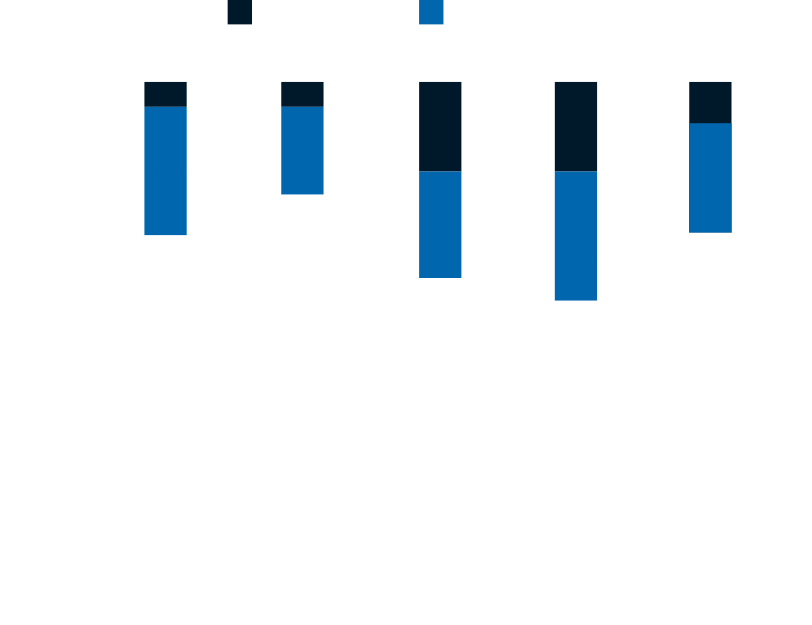

Lifetime risk of developing lung cancer

LUNG CANCER is the second most common cancer in both men and women in the United States. It is the leading cause of cancer death, accounting for about 1 in 5 of all cancer deaths.
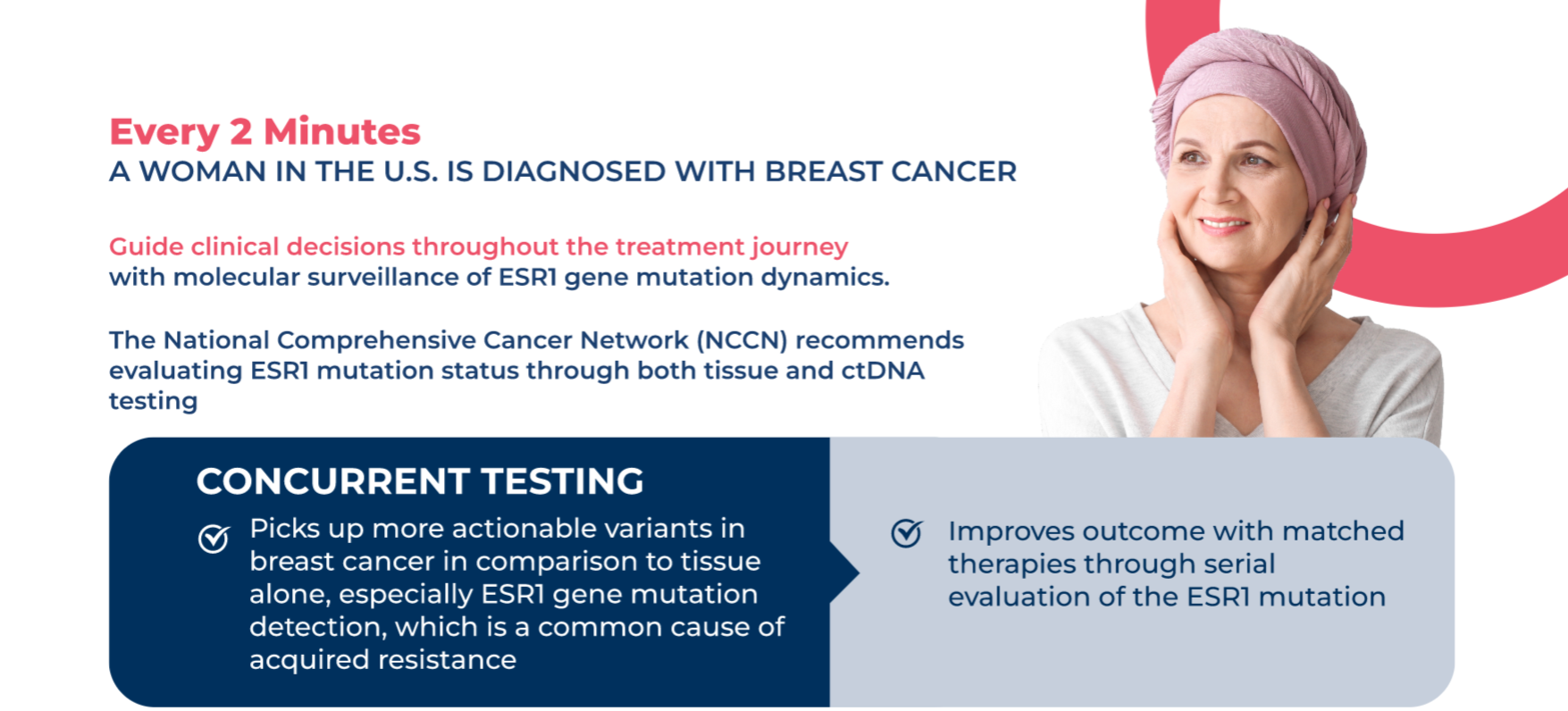
Achieve timely clinical decision making for patients with advanced cancer through
CONCURRENT TESTING
Maximize biomarker detection and identify more clinically actionable variants, such as the EGFR mutation detection
The NCCN Guidelines support the concurrent testing approach
Both liquid ctDNA and tissue testing have very high specificity, complementing each other to reduce turnaround time and increase yield of targetable alteration detection.
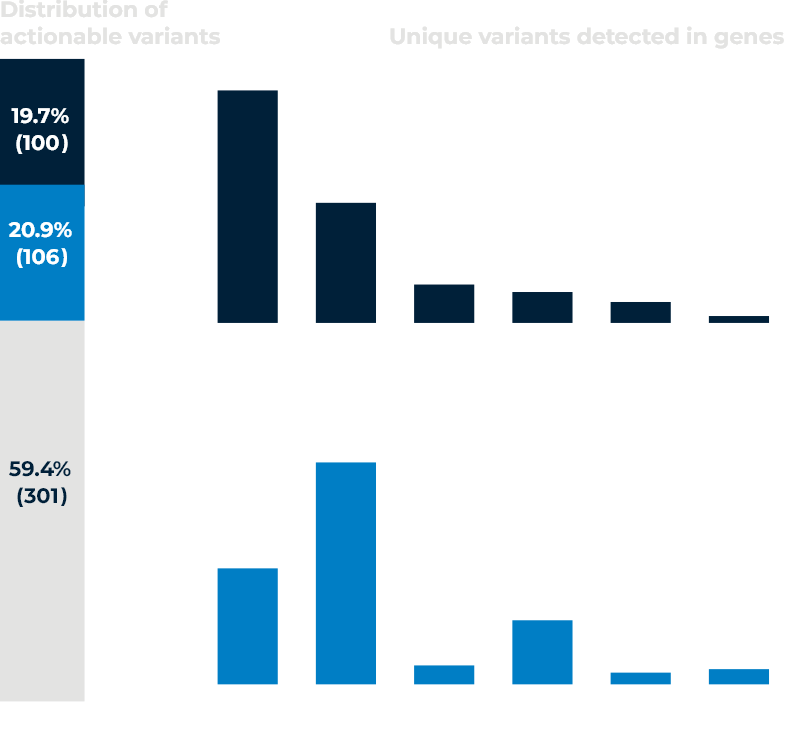
Concurrent Solid and Liquid ctDNA NGS Testing in Metastatic NSCLC patients
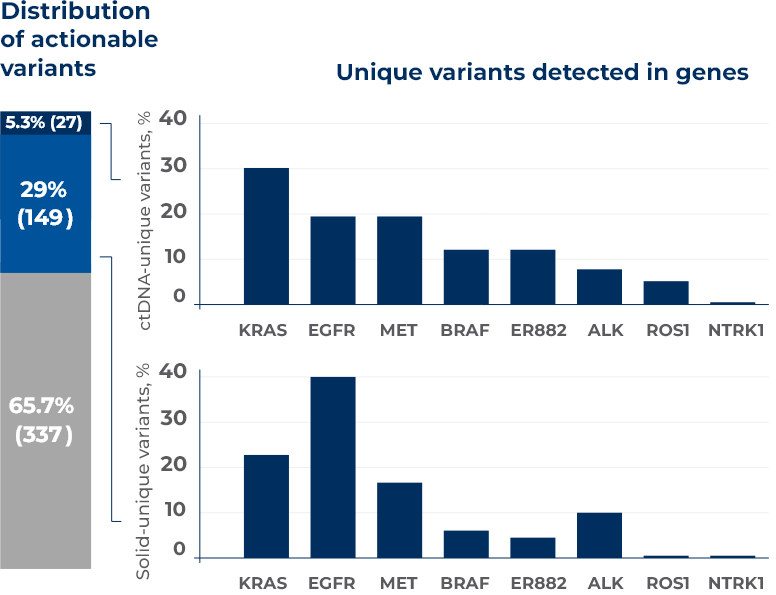 Iams WT, Mackay M, Ben-Shachar R, et al. Concurrent Tissue and Circulating Tumor DNA Molecular Profiling to Detect Guideline-Based Targeted Mutations in a Multicancer Cohort. JAMA Netw Open. 2024;7(1):e2351700. doi:10.1001/jamanetworkopen.2023.51700
Iams WT, Mackay M, Ben-Shachar R, et al. Concurrent Tissue and Circulating Tumor DNA Molecular Profiling to Detect Guideline-Based Targeted Mutations in a Multicancer Cohort. JAMA Netw Open. 2024;7(1):e2351700. doi:10.1001/jamanetworkopen.2023.51700
Reveal your patients' full genomic profile with siParadigm’s Concurrent Testing Options

One Specimen; All the Answers
Comprehensive Precision Oncology Offering
Multiple test modalities, wide
choice of tests



Hassle-free & billing compliant wide “unbundled” choice of testing

Comprehensive profiles ($$$): 500 genes NGS panels (with PARPi, TMB and MSI), FISH and/or IHC tests for advanced or refractory cancers

Focused panels ($$):
<50 gene NGS panels plus FDA-approved
companion diagnostics FISH and/or IHC tests for companion diagnostic testing and prognosis

A la carte tests ($):
Single genes, FISH and/or IHC tests "a la carte" ($): for standard “a la carte” and out of
pocket testing
DeepSight Liquid Biopsy Assays are Analytically Sensitive for Clinically Relevant Genomic Alterations.
NGS Assay Limit of Detection: 0.5% Variant Allele Frequency for Clinically Relevant Single Nucleotide Variants (SNVs) and Insertion-Deletion variants (Indels).




Personalized service: Pathologists’ work and mobile phone numbers on reports for direct access
User-friendly synoptic and fully customizable report to meet your preferences including therapy and clinical trial recommendations, with prioritization of health system’s own trials



Pre and post-test genetic counseling for germline testing

Simplified logistics
(we handle all specimen procurement from the holding facilities, e.g., hospital)

Streamlined workflow integration
(EMR interface or HIPAA-compliant dedicated web portal) for simplified ordering and easy access to results

Professional support for patients and providers
in understanding testing and the implications for clinical care

Free access to molecular pathologist and academic oncologist for test interpretation
Billing We understand the difficulties of the billing process and are committed to making it “hassle-free”.
We require a faxed copy of the completed requisition for pre-authroization. We will inform you with the patient's obligations, if any. Our financial hardship program (see web portal) estimates that >90% of patients will not pay more than their in-network rates

Testimonials
Our customers love us! Read what they have to say below. Aliquam sed justo ligula. Vestibulum nibh erat, pellentesque ut laoreet vitae.
References
Peng G, Lin SY. Exploiting the
homologous recombination DNA repair network for targeted cancer therapy. World J Clin Oncol.
2011 Feb 10;2(2):73-9. doi: 10.5306/wjco.v2.i2.73. PMID: 21603316; PMCID: PMC3095467.
da Cunha Colombo Bonadio RR, Fogace RN,
Miranda VC, Diz MDPE. Homologous recombination deficiency in ovarian cancer: a review of its
epidemiology and management. Clinics (Sao Paulo). 2018 Aug 20;73(suppl 1):e450s. doi:
10.6061/clinics/2018/e450s. PMID: 30133561; PMCID: PMC6096977.
Radovich M, Kiel
PJ, Nance SM, Niland EE, Parsley ME, Ferguson ME, Jiang G, Ammakkanavar NR, Einhorn LH, Cheng
L, Nassiri M, Davidson DD, Rushing DA, Loehrer PJ, Pili R, Hanna N, Callaghan JT, Skaar TC,
Helft PR, Shahda S, O'Neil BH, Schneider BP. Clinical benefit of a precision medicine based
approach for guiding treatment of refractory cancers. Oncotarget. 2016 Aug
30;7(35):56491-56500. doi: 10.18632/oncotarget.10606. PMID: 27447854; PMCID:
PMC5302930.
https://www.fda.gov/drugs/resources-information-approved-drugs/fda-approves-olaparib-germline-brca-mutated-metastatic-breast-cancer
https://www.fda.gov/drugs/drug-approvals-and-databases/fda-approves-talazoparib-gbrcam-her2-negative-locally-advanced-or-metastatic-breast-cancer#:~:text=On%20October%2016%2C%202018%2C%20the,advanced%20or%20metastatic%20breast%20cancer..
https://www.fda.gov/drugs/drug-approvals-and-databases/fda-approves-olaparib-plus-bevacizumab-maintenance-treatment-ovarian-fallopian-tube-or-primary
https://www.fda.gov/drugs/resources-information-approved-drugs/fda-approves-rucaparib-maintenance-treatment-recurrent-ovarian-fallopian-tube-or-primary-peritoneal#:~:text=On%20April%206%2C%202018%2C%20the,partial%20response%20to%20platinum%2Dbased
https://www.fda.gov/drugs/resources-information-approved-drugs/fda-approves-olaparib-gbrcam-metastatic-pancreatic-adenocarcinoma#:~:text=On%20December%2027%2C%202019%2C%20the,an%20FDA%2Dapproved%20test%2C%20whose
https://clinicaltrials.gov/ct2/show/results/NCT02987543
https://www.fda.gov/drugs/drug-approvals-and-databases/fda-approves-olaparib-hrr-gene-mutated-metastatic-castration-resistant-prostate-cancer










